Regulatory networks and molecular mechanisms underlying salt stress tolerance in rice
Wuyun Fang, Ali Raza, Qian Zhu, Qiming Wang, Qun Ren, Mengyang Liu, Shimei Wang, Muhammad Ahmad Hassan
Front Plant Sci.; 2026 Mar 5: 17:1757448. doi: 10.3389/fpls.2026.1757448.
Abstract
Salinity and alkaline stress severely restrict rice productivity by disrupting ionic balance, generating oxidative damage, and impairing growth across developmental stages. Despite the significant advances in the salt tolerance knowledge, rice is very sensitive in contrast to other cereals, which demonstrates gaps in mechanistic understanding and breeding efficiency. This review incorporates the progress in the salt perception, signaling, and stress adaptation, and introduces limitations that slow down the practical improvement. Rice senses salinity using receptor-like kinases and Ca2+-dependent signaling pathway but the initial stages of the response and down-stream phosphorylation cascades have not been characterized well. The reactive oxygen species (ROS) triggered by salinity activate antioxidant mechanisms like AsA-GSH, but it is still not clear how they are spatially and organelle-specifically controlled. Proteomic analyses show extensive reorganization of proteins in signaling, cytoskeleton dynamics, metabolism and protein turnover, but most of the identified candidates have not been validated functionally. Na+ exclusion, vacuolar sequestration, and K+ retention through HKTs, NHXs, and V-ATPases are involved in ion homeostasis, but the interactions between them in tissues have not been fully understood yet. QTL studies have also reported important loci like Saltol and qSKC1 but there are slow advances made in using them in elite cultivars. New multi-omics techniques and CRISPR-based genome editing are currently providing a chance to uncover knowledge gaps. All in all, this review presents an overall framework to develop mechanistic knowledge and speed up breeding salt-resistant varieties of rice.
See https://pubmed.ncbi.nlm.nih.gov/41868506/
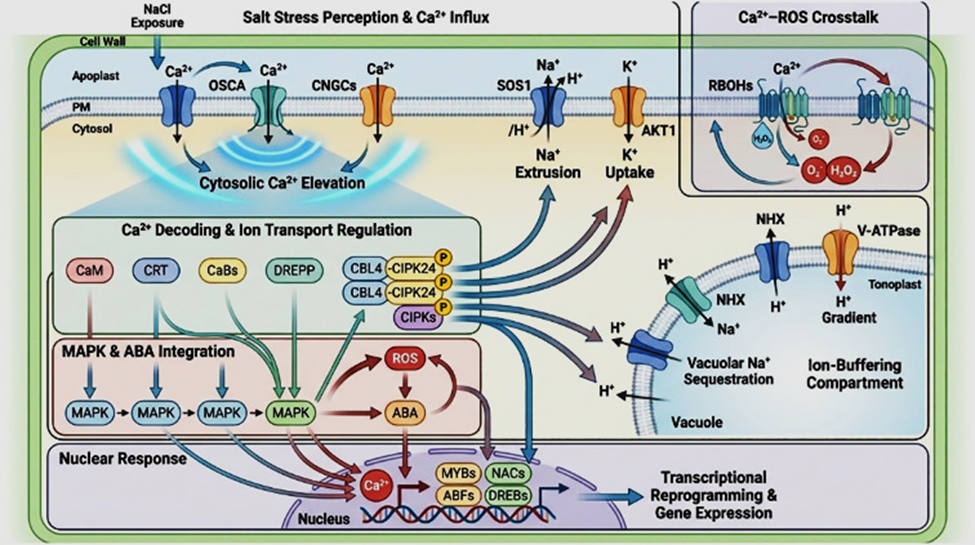
![]() Figure 1: Overview of the major protein groups and pathways modulated in plant roots under salt stress. The root tip is exposed to elevated Na+ and Cl-, and the magnified region summarizes different functional categories showing upregulated (green arrows) or downregulated (red arrows) proteins. Changes are observed in carbohydrate metabolism (ADH, PDC, PGK, APDH, TPI, UGPP, UGDC, MDH, DLDH), protein processing (PPA, TcP1, TEF, RP, PD1), membrane transport (A-ATPase, REM, ANX, V-ATPase), ROS defense (POD, APX, GST, GLYI-II, TRX, SOD), and signaling (CRT, RPK, CaM, STK, DREEP, SR, GEP). Additional changes occur in cell wall polymer dynamics (actin, CCOMT, B23-chain, SAM, COMT, Tuba), transcription-related proteins (RBR, RT, SF, RTP, TrP, DRF), and protein folding components (HSP70, CPN21, CPN60, DnaK-type chaperones). Together these alterations reflect coordinated metabolic, structural, and regulatory adjustments supporting root adaptation to saline conditions.
Figure 1: Overview of the major protein groups and pathways modulated in plant roots under salt stress. The root tip is exposed to elevated Na+ and Cl-, and the magnified region summarizes different functional categories showing upregulated (green arrows) or downregulated (red arrows) proteins. Changes are observed in carbohydrate metabolism (ADH, PDC, PGK, APDH, TPI, UGPP, UGDC, MDH, DLDH), protein processing (PPA, TcP1, TEF, RP, PD1), membrane transport (A-ATPase, REM, ANX, V-ATPase), ROS defense (POD, APX, GST, GLYI-II, TRX, SOD), and signaling (CRT, RPK, CaM, STK, DREEP, SR, GEP). Additional changes occur in cell wall polymer dynamics (actin, CCOMT, B23-chain, SAM, COMT, Tuba), transcription-related proteins (RBR, RT, SF, RTP, TrP, DRF), and protein folding components (HSP70, CPN21, CPN60, DnaK-type chaperones). Together these alterations reflect coordinated metabolic, structural, and regulatory adjustments supporting root adaptation to saline conditions.
Views: 15


