| Engineering broad-spectrum disease-resistant rice by editing multiple susceptibility genes |
|
Rice blast and bacterial blight are important diseases of rice (Oryza sativa) caused by the fungus Magnaporthe oryzae and the bacterium Xanthomonas oryzae pv. oryzae (Xoo), respectively. Breeding rice varieties for broad-spectrum resistance is considered the most effective and sustainable approach to controlling both diseases. Although dominant resistance genes have been extensively used in rice breeding and production, |
|
Hui Tao, Xuetao Shi, Feng He, Dan Wang, Ning Xiao, Hong Fang, Ruyi Wang, Fan Zhang, Min Wang, Aihong Li, Xionglun Liu, Guo-Liang Wang, Yuese Ning Journal of Integrative Plant Biology; First published: 25 June 2021; https://doi.org/10.1111/jipb.13145 ABSTRACTRice blast and bacterial blight are important diseases of rice (Oryza sativa) caused by the fungus Magnaporthe oryzae and the bacterium Xanthomonas oryzae pv. oryzae (Xoo), respectively. Breeding rice varieties for broad-spectrum resistance is considered the most effective and sustainable approach to controlling both diseases. Although dominant resistance genes have been extensively used in rice breeding and production, generating disease-resistant varieties by altering susceptibility (S) genes that facilitate pathogen compatibility remains unexplored. Here, using CRISPR/Cas9 technology, we generated loss-of-function mutants of the S genes Pi21 and Bsr-d1 and showed that they had increased resistance to M. oryzae. We also generated a knockout mutant of the S gene Xa5 that showed increased resistance to Xoo. Remarkably, a triple mutant of all three S genes had significantly enhanced resistance to both M. oryzae and Xoo. Moreover, the triple mutant was comparable to the wild type in regard to key agronomic traits, including plant height, effective panicle number per plant, grain number per panicle, seed setting rate, and thousand-grain weight. These results demonstrate that the simultaneous editing of multiple S genes is a powerful strategy for generating new rice varieties with broad-spectrum resistance.
See: https://onlinelibrary.wiley.com/doi/abs/10.1111/jipb.13145
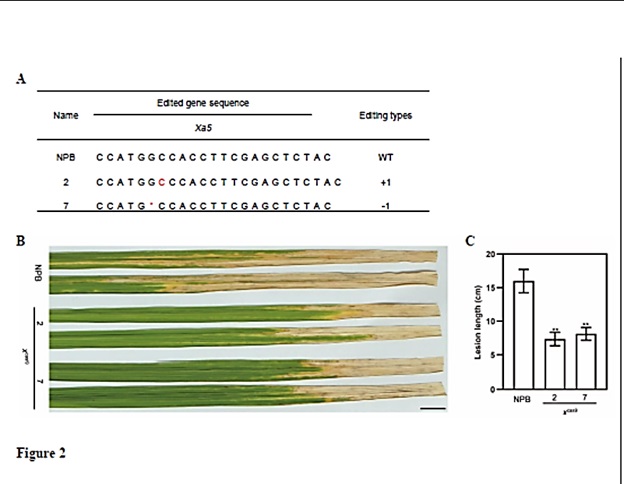
Figure 2. Resistance phenotypes of the xcas9 mutant inoculated with Xoo isolate PXO99A (A) sgRNA sequence and mutations at the target site of Xa5 in the two T0 xcas9 plants. The targeted sequence is from NPB. Red letters ndicate inserted nucleotides. “*” indicates deleted nucleotides. “+” and “” at right indicate insertion and deletion, respectively, and numbers indicate nucleotides. (B) Photograph of infected leaves of NPB and x cas9 mutant plants at 2 weeks after inoculation with PXO99A. Scale bar, 1 cm. (C) Lesion lengths on infected leaves in (B). Values are means ± SE. Asterisks indicate a statistically significant difference, as indicated by Student’s t-test (**P < 0.01). Each experiment had three biological repeats. NPB, Nipponbare; WT, the wild type. |
|
|
|
[ Tin tức liên quan ]___________________________________________________
|


 Curently online :
Curently online :
 Total visitors :
Total visitors :