Comparative Metabolomic Profiling of Resistant and Susceptible Coffea arabica Accessions to Bacterial Pathogen Infection
Salim Makni, Adrian Heckart, Jean-Christophe Cocuron, Lucas Mateus Rivero Rodrigues, Suzete Aparecida Lanza Destéfano, Masako Toma Braghini, Oliveiro Guerreiro Filho, Ana Paula Alonso
Plants (Basel); 2026 Jan 9; 15(2):216. doi: 10.3390/plants15020216.
Abstract
Coffea, a plant species of significant agricultural value used in coffee production, is a key commodity that supports the livelihoods of millions of people worldwide. However, coffee cultivation faces substantial threats from various pathogens, including Pseudomonas coronafaciens pv. garcae (Pcg), the causative agent of bacterial blight. This pathogen compromises coffee plant health, leading to reduced yields and plant death and impacting farmers and large-scale producers. Understanding the mechanisms underlying resistance to Pcg in the leaves of the resistant IAC 2211-6 Coffea arabica accession is crucial for developing effective control strategies. This study aimed to identify candidate biomarkers of resistance by comparing the leaf metabolome of (i) the resistant IAC 2211-6 and the susceptible IAC 125 RN Coffea arabica accessions and (ii) Pcg-infected and uninfected leaves. Untargeted metabolomics revealed distinct metabolic profiles between accessions. Flavonoids were more abundant in susceptible leaves. In contrast, resistant leaves showed increased levels of pipecolic acid ethyl ester, a structural derivative of a key systemic acquired resistance signal, and spiropreussione B, a compound associated with fungal endophytes. These findings highlight candidates potentially linked to resistance and suggest that systemic signaling and beneficial microbial interactions may contribute to resilience.
See https://pubmed.ncbi.nlm.nih.gov/41600023/
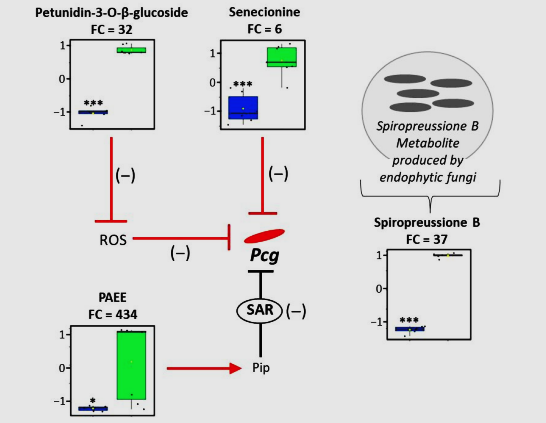
Figure 4. Metabolomic profiling of Coffea leaves indicates distinct metabolite accumulation patterns in the resistant accession. Box plots illustrate the relative abundance of metabolites that accumulate in the resistant (green) versus susceptible (blue) Coffea accessions. These metabolites were integrated into a metabolic and regulatory network. Solid arrows represent single enzymatic reactions, while T-bar symbols indicate inhibitory regulation. The color of arrows and T-bars reflects the degree of confidence for each pathway: black T-bars correspond to systemically acquired resistance (SAR) mechanisms already described in other plant systems, whereas red symbols denote enzymatic reactions or regulations suggested to occur in the resistant Coffea accession. The red arrow specifically marks the putative hydrolysis of pipecolic acid ethyl ester (PAEE) into pipecolic acid (Pip). Statistical significance was assessed using a bilateral Student’s t-test (n = 6, p-value < 0.05: *, p-value < 0.001: ***). Black dots correspond to individual data points. Abbreviation: ROS, reactive oxygen species; Pcg, Pseudomonas coronafaciens pv. garcae.
Views: 40


