|
Orkun Çoruh, Anna Frank, Hideaki Tanaka, Akihiro Kawamoto, Eithar El-Mohsnawy, Takayuki Kato, Keiichi Namba, Christoph Gerle, Marc M. Nowaczyk & Genji Kurisu
Communications Biology volume 4, Article number: 304 (8 March 2021)
Abstract
A high-resolution structure of trimeric cyanobacterial Photosystem I (PSI) from Thermosynechococcus elongatus was reported as the first atomic model of PSI almost 20 years ago. However, the monomeric PSI structure has not yet been reported despite long-standing interest in its structure and extensive spectroscopic characterization of the loss of red chlorophylls upon monomerization. Here, we describe the structure of monomeric PSI from Thermosynechococcus elongatus BP-1. Comparison with the trimer structure gave detailed insights into monomerization-induced changes in both the central trimerization domain and the peripheral regions of the complex. Monomerization-induced loss of red chlorophylls is assigned to a cluster of chlorophylls adjacent to PsaX. Based on our findings, we propose a role of PsaX in the stabilization of red chlorophylls and that lipids of the surrounding membrane present a major source of thermal energy for uphill excitation energy transfer from red chlorophylls to P700.
See https://www.nature.com/articles/s42003-021-01808-9
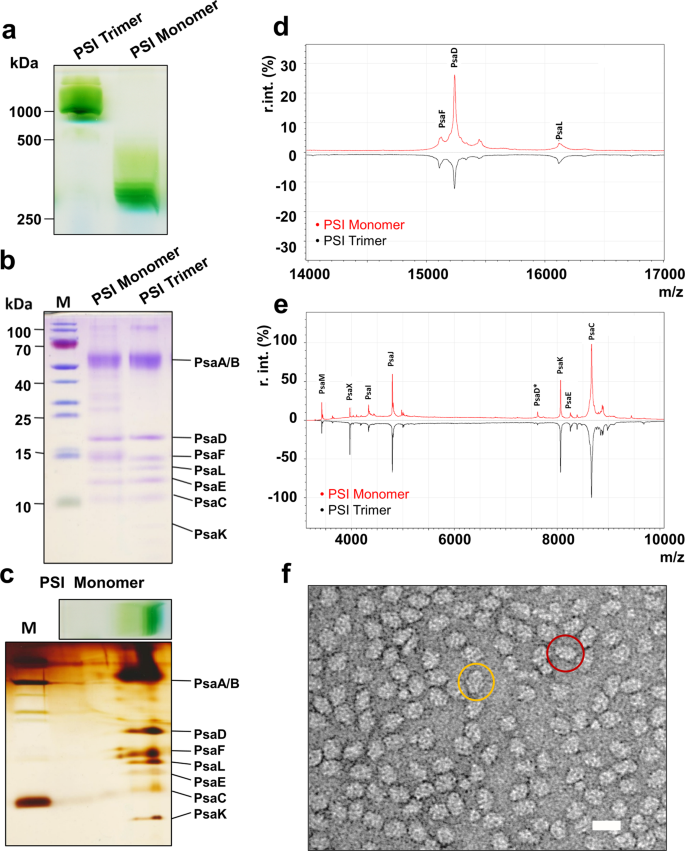
Figure 1: Biochemical characterization of the purified PSI monomer complex from T. elongatus.
a Blue-native (BN) PAGE of the PSI monomer next to a PSI trimer control (3.5 µg Chl/lane). b SDS-PAGE of monomeric and trimeric PSI in comparison (2.5 µg Chl/lane) c Two-dimensional (2D)-PAGE of monomeric PSI (3.5 µg Chl/BN-lane). d, e MALDI-TO Fmass-spectrometry of monomeric and trimeric PSI complexes, mass range: d 14,000–17,000 m/z and e 3000–10,000 m/z (see also Table 1), *z=2. f Negative stain EM image of PSI monomer with a top view encircled in yellow and a side view encircled in red. Scale bar corresponds to 20 nm.
|
[ Other News ]___________________________________________________
|