|
A Genome-Wide Association Study of Field Resistance to Magnaporthe oryzae in Rice
Wednesday, 2016/09/07 | 08:39:57
|
|
Zhu D, Kang H, Li Z, Liu M, Zhu X, Wang Y, Wang D, Wang Z, Liu W, Wang GL.
Rice (N Y). 2016 Dec;9(1):44. doi: 10.1186/s12284-016-0116-3. Epub 2016 Aug 30. AbstractBACKGROUND:Breeding of rice cultivars with long-lasting resistance to the rice blast fungus Magnaporthe oryzae is difficult, and identification of new resistance genes is essential. Most of the loci associated with blast resistance against M. oryzae in rice have been identified in controlled environments and with single isolates, and such loci may confer resistance to only a small faction of the M. oryzae strains. In the field, however, rice is commonly attacked by multiple strains. Research is therefore needed to identify loci that confer resistance in the field, i.e., "field blast resistance". To identify loci associated with field blast resistance (LAFBRs), we conducted a genome-wide association study (GWAS) using the rice diversity panel 1 (RDP1) cultivars. These cultivars were evaluated in the field in three major rice production areas of China. RESULTS:GWAS identified 16 LAFBRs. Among them, 13 are novel and the other three are co-localized with known blast resistance regions. Seventy-four candidate genes are identified in the 16 LAFBR regions, which encode receptor-like protein kinases, transcription factors, and other defense-related proteins. Using the rice transcriptome data, compared with the rice-rice blast compatible interaction, we identified seven candidate genes that are significantly up-regulated and five genes that are significantly down-regulated in the incompatible interaction among the candidate genes. CONCLUSIONS:We identified 16 LAFBRs involved in field resistance to M. oryzae and 20 cultivars that exhibit high levels of resistance in both the field and growth chamber. The resistant cultivars and the SNP markers identified in this study should be useful for marker-assisted selection of new rice cultivars that confer high levels of resistance against M. oryzae field populations.
See: http://www.ncbi.nlm.nih.gov/pubmed/27576685
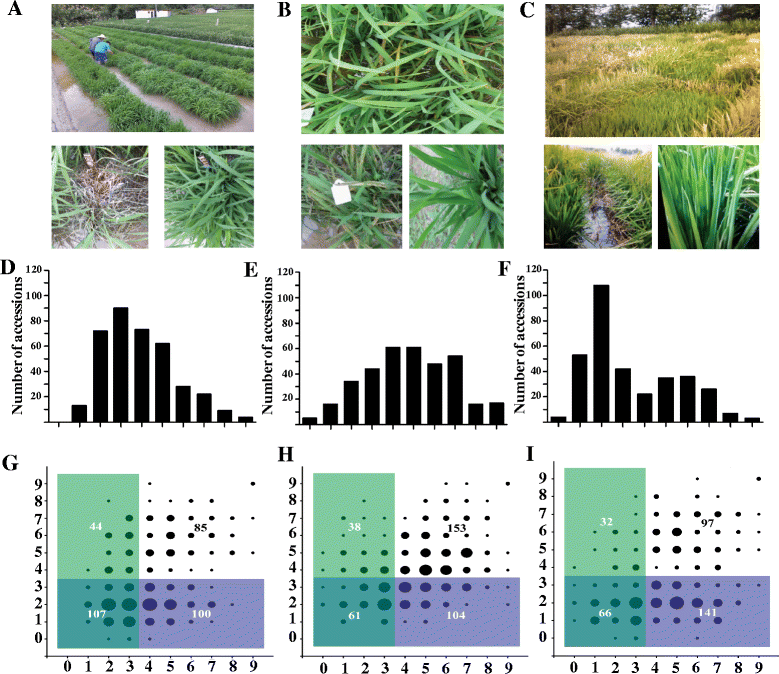
Fig. 1. Field blast evaluation of the RDP1 cultivars and disease resistance scores of the cultivars in the three locations. a–c Seedlings of RDP1 in Shanghang (Fujian province), Taojiang (Hunan province) and Wuchang (Heilongjiang province) blast nurseries, respectively. d–f Distribution of blast disease resistance scores of RDP1 in Shanghang, Taojiang and Wuchang respectively. X axis represents the disease scales, Y-axis represents the number of cultivars. g Pair-wise comparison of disease resistance between Shanghang (X-axis) and Wuchang (Y-axis). h Pair-wise comparison of disease resistance between Taojiang (X-axis) and Shanghang (Y-axis). i Pair-wise comparison of disease resistance between Taojiang (X-axis) and Wuchang (Y-axis). The area of black circles represents the accession numbers. The overlap region with double colors represents the resistant cultivars (0–3) at both locations and the single color region represents the resistant accessions (0–3) at one location but susceptible (4–9) at the other location. |
|
|
|
[ Other News ]___________________________________________________
|


 Curently online :
Curently online :
 Total visitors :
Total visitors :