|
Identification of Coilin Mutants in a Screen for Enhanced Expression of an Alternatively Spliced GFP Reporter Gene in Arabidopsis thaliana
Tuesday, 2016/08/16 | 07:56:23
|
|
Tatsuo Kanno, Wen-Dar Lin, Jason L. Fu, Ming-Tsung Wu, Ho-Wen Yang, Shih-Shun Lin, Antonius J. M. Matzke, and Marjori Matzke Genetics August 2016 203 (4) 1709-1720 AbstractCoilin is a marker protein for subnuclear organelles known as Cajal bodies, which are sites of various RNA metabolic processes including the biogenesis of spliceosomal small nuclear ribonucleoprotein particles. Through self-associations and interactions with other proteins and RNA, coilin provides a structural scaffold for Cajal body formation. However, despite a conspicuous presence in Cajal bodies, most coilin is dispersed in the nucleoplasm and expressed in cell types that lack these organelles. The molecular function of coilin, particularly of the substantial nucleoplasmic fraction, remains uncertain. We identified coilin loss-of-function mutations in a genetic screen for mutants showing either reduced or enhanced expression of an alternatively spliced GFP reporter gene in Arabidopsis thaliana. The coilin mutants feature enhanced GFP fluorescence and diminished Cajal bodies compared with wild-type plants. The amount of GFP protein is several-fold higher in the coilin mutants owing to elevated GFP transcript levels and more efficient splicing to produce a translatable GFP mRNA. Genome-wide RNA-sequencing data from two distinct coilin mutants revealed a small, shared subset of differentially expressed genes, many encoding stress-related proteins, and, unexpectedly, a trend toward increased splicing efficiency. These results suggest that coilin attenuates splicing and modulates transcription of a select group of genes. The transcriptional and splicing changes observed in coilin mutants are not accompanied by gross phenotypic abnormalities or dramatically altered stress responses, supporting a role for coilin in fine tuning gene expression. Our GFP reporter gene provides a sensitive monitor of coilin activity that will facilitate further investigations into the functions of this enigmatic protein.
See http://www.genetics.org/content/203/4/1709?etoc
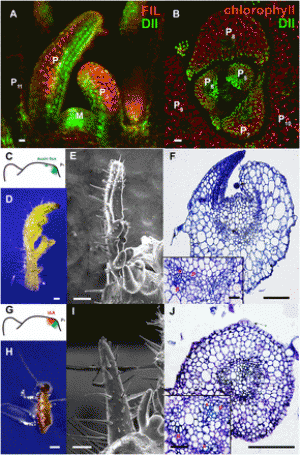
Figure 1 GFP reporter gene system and GFP phenotypes in mutants. (A) The GFP reporter gene in the T line and alternative splicing of GFP pre-mRNA. The GFP coding region (green bar) is under the control of virus-derived transcriptional regulatory elements: a truncated 35S promoter (TATA) and the endogenous pararetrovirus (EPRV) enhancer (∼1.2 kb, black bar), which contains a tandem repeat (three copies of a 41–42-bp monomer, arrowheads) in the 5′ distal portion. Transcription of GPF pre-mRNA begins around the tandem repeat region. Two opposing arrows above the diagram indicate the positions of primers used for RT-PCR to detect three major GFP transcripts: the untranslatable “long” and “middle” transcripts, and a “short” translatable mRNA. The middle and short transcripts result from alternative splicing of U2-type introns containing canonical (GT-AG) and noncanonical (AT-AC) splice sites, respectively (Sasaki et al., 2015). (B) GFP phenotypes in seedlings. The wild-type T line shows an intermediate level of GFP fluorescence visible mainly in the stem (hypocotyl) and shoot and root apices of young seedlings. Mutants obtained following EMS mutagenesis of the T line could display either reduced (“Weak”) or stronger (“Hyper”) GFP fluorescence. Cotyledons (the first set of leaves sprouting from the seed) appear red owing to auto-fluorescence of chlorophyll at the excitation wavelength for GFP. |
|
|
|
[ Other News ]___________________________________________________
|


 Curently online :
Curently online :
 Total visitors :
Total visitors :
(25).png)