|
Arabidopsis ROCK1 transports UDP-GlcNAc/UDP-GalNAc and regulates ER protein quality control and cytokinin activity
Monday, 2015/01/12 | 08:31:03
|
|
Michael C. E. Niemanna, Isabel Bartrinaa, Angel Ashikovb, Henriette Webera, Ondřej Novákc, Lukáš Spíchalc, Miroslav Strnadc, Richard Strasserd, Hans Bakkerb, Thomas Schmüllinga, and Tomáš Wernera,1 Significance
Nucleotide sugars are donor substrates for the formation of glycan modifications, which are important for the function of many macromolecules such as proteins and lipids. Although most of the glycosylation reactions occur in the endoplasmic reticulum (ER) and Golgi of eukaryotic cells, nucleotide sugar activation occurs in the cytosol and specific transporters must carry these molecules across the membrane. We identified REPRESSOR OF CYTOKININ DEFICIENCY 1 (ROCK1) as an ER-localized transporter of UDP-GlcNAc and UDP-GalNAc in plants. In contrast to animals, nothing is known about the function of the two respective sugar residues in the plant ER. We demonstrate that ROCK1-mediated transport plays a role in the ER-associated protein quality control and loss of ROCK1 enhances cytokinin responses by suppressing the activity of cytokinin-degrading CKX proteins. Abstract
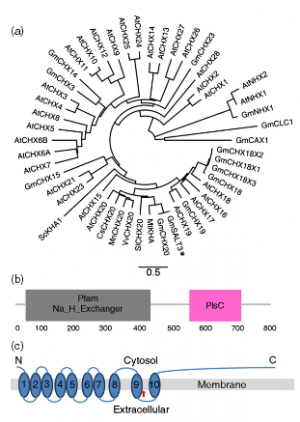
The formation of glycoconjugates depends on nucleotide sugars, which serve as donor substrates for glycosyltransferases in the lumen of Golgi vesicles and the endoplasmic reticulum (ER). Import of nucleotide sugars from the cytosol is an important prerequisite for these reactions and is mediated by nucleotide sugar transporters. Here, we report the identification of REPRESSOR OF CYTOKININ DEFICIENCY 1 (ROCK1, At5g65000) as an ER-localized facilitator of UDP-N-acetylglucosamine (UDP-GlcNAc) and UDP-N-acetylgalactosamine (UDP-GalNAc) transport in Arabidopsis thaliana. Mutant alleles of ROCK1 suppress phenotypes inferred by a reduced concentration of the plant hormone cytokinin. This suppression is caused by the loss of activity of cytokinin-degrading enzymes, cytokinin oxidases/dehydrogenases (CKXs). Cytokinin plays an essential role in regulating shoot apical meristem (SAM) activity and shoot architecture. We show that rock1 enhances SAM activity and organ formation rate, demonstrating an important role of ROCK1 in regulating the cytokinin signal in the meristematic cells through modulating activity of CKX proteins. Intriguingly, genetic and molecular analysis indicated that N-glycosylation of CKX1 was not affected by the lack of ROCK1-mediated supply of UDP-GlcNAc. In contrast, we show that CKX1 stability is regulated in a proteasome-dependent manner and that ROCK1 regulates the CKX1 level. The increased unfolded protein response in rock1 plants and suppression of phenotypes caused by the defective brassinosteroid receptor bri1-9 strongly suggest that the ROCK1 activity is an important part of the ER quality control system, which determines the fate of aberrant proteins in the secretory pathway.
See: http://www.pnas.org/content/112/1/291.abstract.html?etoc PNAS Janurary 6, 2014; Vol.112, no.1: 291-296
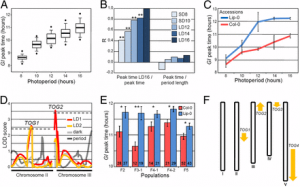
Fig. 1. rock1 suppresses the cytokinin deficiency phenotype by repressing CKX activity. (A) Suppression of the 35S:CKX1 shoot phenotype by rock1-1 mutation in 4-wk-old plants. (B) Relative transcript abundance of A-type ARR genes in shoots of soil-grown seedlings 10 d after germination (dag) measured by quantitative real-time PCR. Data are means ± SD (n = 4; *P < 0.05, t test). (C) Effect of rock1-1 on shoot development in plants expressing 35S:CKX2 or 35S:CKX3. The shoot fresh weight of soil-grown plants was determined 17 dag (means ± SD, n ≥ 15). Significant differences to wild type were determined by t test (*P < 0.05). (D) CKX activity measured in total protein extracts. Activity is expressed relative to wild type. Values are means ± SD (n ≥ 3). Significant differences to the respective CKX overexpression line were determined by t test (*P < 0.05).
|
|
|
|
[ Other News ]___________________________________________________
|


 Curently online :
Curently online :
 Total visitors :
Total visitors :
(14).png)