|
Auxin depletion from leaf primordia contributes to organ patterning
Friday, 2015/01/02 | 17:30:53
|
|
Jiyan Qi, Ying Wang, Ting Yu, Alexandre Cunha, Binbin Wu, Teva Vernoux, Elliot Meyerowitz, and Yuling Jiao SignificanceStem cells not only initiate organs, but may also contribute to organ patterning, at least in the shoot apex of flowering plants: classical microsurgical experiments imply that the shoot apical meristem promotes development of the leaf adaxial side, i.e., the upper side. In this study, we show the existence of a transient low auxin zone in the adaxial side that contributes to adaxial development. We further find that this adaxial low auxin zone results from auxin transport from leaves to the shoot apex. Thus, it is not a positive signal from stem cells, but departure of a signaling molecule from primordia to stem cells, that delivers polarity information—opposite to what is generally assumed. AbstractStem cells are responsible for organogenesis, but it is largely unknown whether and how information from stem cells acts to direct organ patterning after organ primordia are formed. It has long been proposed that the stem cells at the plant shoot apex produce a signal, which promotes leaf adaxial-abaxial (dorsoventral) patterning. Here we show the existence of a transient low auxin zone in the adaxial domain of early leaf primordia. We also demonstrate that this adaxial low auxin domain contributes to leaf adaxial-abaxial patterning. The auxin signal is mediated by the auxin-responsive transcription factor MONOPTEROS (MP), whose constitutive activation in the adaxial domain promotes abaxial cell fate. Furthermore, we show that auxin flow from emerging leaf primordia to the shoot apical meristem establishes the low auxin zone, and that this auxin flow contributes to leaf polarity. Our results provide an explanation for the hypothetical meristem-derived leaf polarity signal. Opposite to the original proposal, instead of a signal derived from the meristem, we show that a signaling molecule is departing from the primordium to the meristem to promote robustness in leaf patterning.
See http://www.pnas.org/content/111/52/18769.abstract.html?etoc PNAS December 30, 2014 vol. 111 no. 52 18769–18774
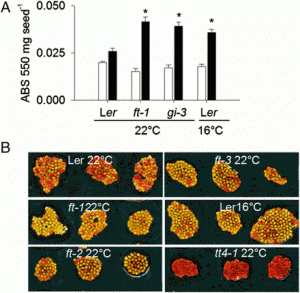
Fig. 1. Transient adaxial low auxin domain is important for leaf polarity patterning. (A and B) Longitudinal and transverse sections through Arabidopsis SAM and leaf primordia region. DII-Venus signals are shown in green in A and B, and chlorophyll autofluorescence is in red (B). DII-Venus signal is enriched in the adaxial domain from P2 to P9. The abaxial domain in A has pFIL::DsRed-N7 (red) expression. Stronger DII-Venus signals in the boundary and adjacent adaxial domain indicated weaker auxin signaling input. Images in Fig. S1 show DII-Venus signals in leaf primordia of additional stages. (C–F) Control tomato leaf primordia, showing schematic diagram of the meristem surface auxin flux direction (C), early primordium 4–5 d after emergence (D), 7 d after emergence (E), and a transverse section through the midrib and adjacent laminal regions with close-up insertion of vascular strand (F). Note that phloem cells (p) surround the xylem (x) elements. (G–J), Tomato leaf primordia after adaxial IAA microapplication, showing site of microapplication (G), early leaf (H), and more mature (I) primordia with strong defects in adaxial-abaxial polarity, and a transverse section through the midrib and adjacent laminal regions (J). More images are shown in Figs. S5 and S6. (Scale bars: A and B, 20 μm; C–J, 200 μm; F and J Inset, 50 μm.) |
|
|
|
[ Other News ]___________________________________________________
|


 Curently online :
Curently online :
 Total visitors :
Total visitors :
.gif)