|
The Blast Fungus Decoded: Genomes in Flux.
Sunday, 2018/04/29 | 07:20:25
|
|
Langner T, Białas A, Kamoun S. MBio. 2018 Apr 17;9(2). pii: e00571-18. doi: 10.1128/mBio.00571-18. AbstractPlant disease outbreaks caused by fungi are a chronic threat to global food security. A prime case is blast disease, which is caused by the ascomycete fungus Magnaporthe oryzae (syn. Pyricularia oryzae), which is infamous as the most destructive disease of the staple crop rice. However, despite its Linnaean binomial name, M. oryzae is a multihost pathogen that infects more than 50 species of grasses. A timely study by P. Gladieux and colleagues (mBio 9:e01219-17, 2018, https://doi.org/10.1128/mBio.01219-17) reports the most extensive population genomic analysis of the blast fungus thus far. M. oryzae consists of an assemblage of differentiated lineages that tend to be associated with particular host genera. Nonetheless, there is clear evidence of gene flow between lineages consistent with maintaining M. oryzae as a single species. Here, we discuss these findings with an emphasis on the ecologic and genetic mechanisms underpinning gene flow. This work also bears practical implications for diagnostics, surveillance, and management of blast diseases.
See: http://mbio.asm.org/content/9/2/e00571-18/F1.expansion.html
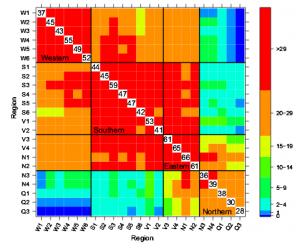
Figure 1: Coinfection on common host plants may facilitate genetic exchange between distinct Magnaporthe oryaze lineages. (A) Although the M. oryzae species complex has the capacity to infect a wide range of grass species, individual lineages tend to be host specific and infect only a limited number of hosts, e.g., lineage 1 (yellow) and lineage 2 (brown) infect host 1 (light green) and host 2 (green), respectively. However, under certain ecologic conditions, two distinct lineages may occasionally colonize or jump on a common host (dark green). Coinfection on a shared host enables interlineage genetic exchange, such as recombination and gene flow, leaving footprints in pathogen genomes. Subsequent selection pressure imposed by the original hosts can either counterselect introgressed segments (lineage 2) or lead to the emergence of a new (sub)lineage (lineage 1*). (B) Similarity plot between different M. oryzae lineages with a genomic region from lineage 2 introgressed into lineage 1, leading to the emergence of lineage 1* (gray box). |
|
|
|
[ Other News ]___________________________________________________
|


 Curently online :
Curently online :
 Total visitors :
Total visitors :
(43).png)