|
The energy sensor OsSnRK1a confers broad-spectrum disease resistance in rice.
Thursday, 2018/03/08 | 07:42:27
|
|
Filipe O, De Vleesschauwer D, Haeck A, Demeestere K, Höfte M.
Sci Rep. 2018 Mar 1;8(1):3864. doi: 10.1038/s41598-018-22101-6. AbstractSucrose non-fermenting-1-related protein kinase-1 (SnRK1) belongs to a family of evolutionary conserved kinases with orthologs in all eukaryotes, ranging from yeasts (SnF1) to mammals (AMP-Activated kinase). These kinases sense energy deficits caused by nutrient limitation or stress and coordinate the required adaptations to maintain energy homeostasis and survival. In plants, SnRK1 is a global regulator of plant metabolism and is also involved in abiotic stress responses. Its role in the response to biotic stress, however, is only starting to be uncovered. Here we studied the effect of altered SnRK1a expression on growth and plant defense in rice. OsSnRK1a overexpression interfered with normal growth and development and increased resistance against both (hemi)biotrophic and necrotrophic pathogens, while OsSnRK1a silencing in RNAi lines increased susceptibility. OsSnRK1a overexpression positively affected the salicylic acid pathway and boosted the jasmonate-mediated defense response after inoculation with the blast fungus Pyricularia oryzae. Together these findings strongly suggest OsSnRK1a to be involved in plant basal immunity and favor a model whereby OsSnRK1a acts as a master switch that regulates growth-immunity trade-offs.
See: https://www.nature.com/articles/s41598-018-22101-6
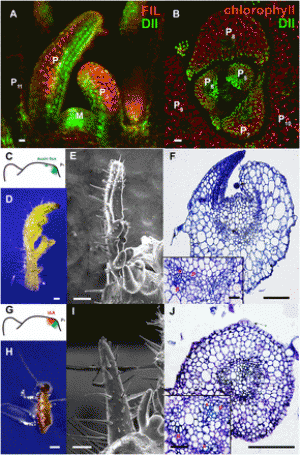
Figure 1: OsSnRK1a overexpression negatively affects plant growth and development. (a) Diagrammatic representation of the rice Sucrose Non-fermenting-1 related kinase 1 A (OsSnRK1a). Protein motif and domains of OsSnRK1a were identified using the Simple Architecture Research Tool (SMART) (http://smart.embl-heidelberg.de) and NCBI-CDD. (http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi). S_TKc, Serine/Threonine protein kinase catalytic domain; UBA, ubiquitin-associated domain; KA1, kinase-associated 1 motif. (b) to (f) Phenotypes of T1 OsSnRK1a overexpressing transgenic rice plants. Azygous controls = null segregants (NS, progeny of lines OsSnRK1a-OX 13–2, 17–4) and Kitaake wild type (WT). OsSnRK1a-OX = progeny of OsSnRK1a-overexpressing lines 5-1, 5-3, 9-5, 10-5, 14-2, and 14-8. When 12 weeks old, plants were scored for several growth parameters. Overexpression of OsSnRK1a inhibited (b) plant growth, (c) tiller number, (d) seed yield, (e) epidermal and mesophylic cell elongation, and (f) root growth. Box plots in (b) to (d) show box and whiskers (Min to Max), open circles represent outliers outside the range of the whiskers. Welsh’s t-test for (b), (c) and (d) revealed significant results with p = 0.001621, p = 0.0005032, and p = 0.001197, respectively. Pictures are representative of the lines in bold. Bars in (b), (e) and (f) are 7 cm, 50 μm and 3 cm respectively. Pictures in (d) show seed yield of a single representative plant at week 13. Asterisks in (e) represent significant differences compared to Kitaake plants (Mann-Whitney test, ns = p > 0.05, *p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001). |
|
|
|
[ Other News ]___________________________________________________
|


 Curently online :
Curently online :
 Total visitors :
Total visitors :
(47).png)