|
Deciphering the genetic basis of resistance to soybean cyst nematode combining IBD and association mapping
Tuesday, 2023/03/21 | 08:16:01
|
|
Yu Tian, Delin Li, Xueqing Wang, Hao Zhang, Jiajun Wang, Lijie Yu, Changhong Guo, Xiaoyan Luan, Xinlei Liu, Hongjie Li, Jochen C. Reif, Ying-hui Li & Li-juan Qiu Theoretical and Applied Genetics March 2023; vol. 136, Article number: 50 (2023) Published: 13 March 2023 Key messageIBD analysis clarified the dynamics of chromosomal recombination during the ZP pedigree breeding process and identified ten genomic regions resistant to SCN race3 combining association mapping. AbstractSoybean cyst nematode (SCN, Heterodera glycines Ichinohe) is one of the most devastating pathogens for soybean production worldwide. The cultivar Zhongpin03-5373 (ZP), derived from SCN-resistant progenitor parents, Peking, PI 437654 and Huipizhi Heidou, is an elite line with high resistance to SCN race3. In the current study, a pedigree variation map was generated for ZP and its ten progenitors using 3,025,264 high-quality SNPs identified from an average of 16.2 × re-sequencing for each genome. Through identity by decent (IBD) tracking, we showed the dynamic change of genome and detected important IBD fragments, which revealed the comprehensively artificial selection of important traits during ZP breeding process. A total of 2,353 IBD fragments related to SCN resistance including SCN-resistant genes rhg1, rhg4 and NSFRAN07 were identified based on the resistant-related genetic paths. Moreover, 23 genomic regions underlying resistance to SCN race3 were identified by genome-wide association study (GWAS) in 481 re-sequenced cultivated soybeans. Ten common loci were found by both IBD tracking and GWAS analysis. Haplotype analysis of 16 potential candidate genes suggested a causative SNP (C/T, − 1065) located in the promoter of Glyma.08G096500 and encoding a predicted TIFY5b-related protein on chr8 was highly correlated with SCN race3 resistance. Our results more thoroughly elucidated the dynamics of genomic fragments during ZP pedigree breeding and the genetic basis of SCN resistance, which will provide useful information for gene cloning and the development of resistant soybean cultivars using a marker-assisted selection approach.
See https://link.springer.com/article/10.1007/s00122-023-04268-3
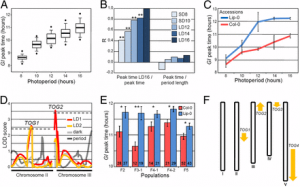
Figure 4: Identification of genetic loci controlling resistance to SCN race3 based on genome-wide association analysis. A Whole genome-wide Manhattan plot. Yellow dots indicate the SNPs significantly associated with SCN races resistance (FDR, P < 0.01). B Regional Manhattan plot (top) and LD heatmap (bottom) for locus SCN3-7–1. The corresponding annotated genes located in the regional plot are drawn in black arrows in the middle part. The bottom part shows the LD block of region plot. C The difference of FI between two allele chr7:15,970,845 genotypes (GG/AA) in soybean accessions. P value was determined by one-way ANOVA analysis. D Geographical distribution of SNP chr7:15,970,845 genotypes
|
|
|
|
[ Other News ]___________________________________________________
|


 Curently online :
Curently online :
 Total visitors :
Total visitors :
(232).png)