|
Photosynthesis in rice is increased by CRISPR/Cas9-mediated transformation of two truncated light-harvesting antenna
Tuesday, 2023/02/14 | 08:12:21
|
|
Daniel Caddell, Noah J. Langenfeld , Madigan JH. Eckels, Shuyang Zhen, Rachel Klaras, Laxmi Mishra, Bruce Bugbee and Devin Coleman-Derr
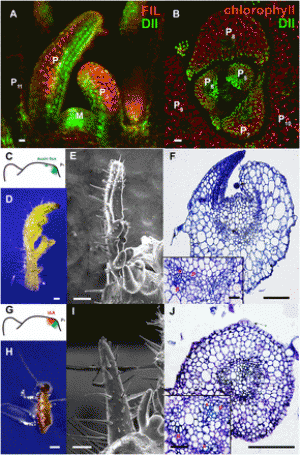
Front. Plant Sci., 19 January 2023; Sec. Photosynthesis and Photobiology ABSTRACTPlants compete for light partly by over-producing chlorophyll in leaves. The resulting high light absorption is an effective strategy for out competing neighbors in mixed communities, but it prevents light transmission to lower leaves and limits photosynthesis in dense agricultural canopies. We used a CRISPR/Cas9-mediated approach to engineer rice plants with truncated light-harvesting antenna (TLA) via knockout mutations to individual antenna assembly component genes CpSRP43, CpSRP54a, and its paralog, CpSRP54b. We compared the photosynthetic contributions of these components in rice by studying the growth rates of whole plants, quantum yield of photosynthesis, chlorophyll density and distribution, and phenotypic abnormalities. Additionally, we investigated a Poales-specific duplication of CpSRP54. The Poales are an important family that includes staple crops such as rice, wheat, corn, millet, and sorghum. Mutations in any of these three genes involved in antenna assembly decreased chlorophyll content and light absorption and increased photosynthesis per photon absorbed (quantum yield). These results have significant implications for the improvement of high leaf-area-index crop monocultures.
See https://www.frontiersin.org/articles/10.3389/fpls.2023.1050483/full
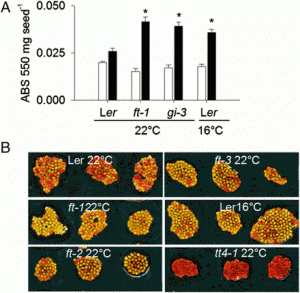
FIGURE 1: TLA3 mutants have reduced height and chlorophyll content. (A) Measurements of plant height for three independent CRISPR/Cas9-generated TLA3 mutant lines as compared to Kitaake wild type (WT). (B) Measurements of chlorophyll content for these three mutants, as measured using a SPAD meter. (C) Representative images of the homozygous parental line of each of the three mutant lineages characterized in (A) and (B), demonstrating reduced stature and a pale green leaf phenotype. (D) Protein domain model of validated CRISPR/Cas9-mediated TLA3 mutations generated using two sgRNAs indicating the relative position and size in base pairs of large deletions within CpSRP43. Frameshift mutations occurred in the middle of chromodomain 1 in each line. CTP, Chloroplast targeting peptide.
|
|
|
|
[ Other News ]___________________________________________________
|


 Curently online :
Curently online :
 Total visitors :
Total visitors :
(205).png)