|
Premeiotic 24-nt phasiRNAs are present in the Zea genus and unique in biogenesis mechanism and molecular function
Monday, 2024/05/27 | 08:12:12
|
|
Junpeng Zhan, Sébastien Bélanger, Scott Lewis, Chong Teng, Madison McGregor, Aleksandra Beric, Michael A. Schon, Michael D. Nodine, and Blake C. Meyers
PNAS May 13, 2024; 121 (21) e2402285121 SignificanceWe previously reported two classes of reproductive phasiRNAs (phased, small interfering RNAs) in maize, the premeiotic 21-nt (nucleotides) phasiRNAs and the meiotic 24-nt phasiRNAs. Here, we report a third class of reproductive phasiRNAs—premeiotic 24-nt phasiRNAs—that are present in the Zea genus, including all five maize inbred lines and three teosinte species/subspecies that we examined, plus rice. We show that in the Zea genus, the premeiotic 24-nt phasiRNAs are distinct from the meiotic 24-nt phasiRNAs in triggering mechanism, effector protein, and molecular function. AbstractReproductive phasiRNAs (phased, small interfering RNAs) are broadly present in angiosperms and play crucial roles in sustaining male fertility. While the premeiotic 21-nt (nucleotides) phasiRNAs and meiotic 24-nt phasiRNA pathways have been extensively studied in maize (Zea mays) and rice (Oryza sativa), a third putative category of reproductive phasiRNAs–named premeiotic 24-nt phasiRNAs–have recently been reported in barley (Hordeum vulgare) and wheat (Triticum aestivum). To determine whether premeiotic 24-nt phasiRNAs are also present in maize and related species and begin to characterize their biogenesis and function, we performed a comparative transcriptome and degradome analysis of premeiotic and meiotic anthers from five maize inbred lines and three teosinte species/subspecies. Our data indicate that a substantial subset of the 24-nt phasiRNA loci in maize and teosinte are already highly expressed at the premeiotic phase. The premeiotic 24-nt phasiRNAs are similar to meiotic 24-nt phasiRNAs in genomic origin and dependence on DCL5 (Dicer-like 5) for biogenesis, however, premeiotic 24-nt phasiRNAs are unique in that they are likely i) not triggered by microRNAs, ii) not loaded by AGO18 proteins, and iii) not capable of mediating PHAS precursor cleavage. In addition, we also observed a group of premeiotic 24-nt phasiRNAs in rice using previously published data. Together, our results indicate that the premeiotic 24-nt phasiRNAs constitute a unique class of reproductive phasiRNAs and are present more broadly in the grass family (Poaceae) than previously known.
See https://www.pnas.org/doi/10.1073/pnas.2402285121
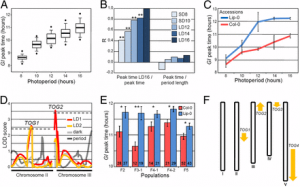
Figure 4: Total sRNA-seq and TraPR sRNA-seq analyses of the ago18 triple mutant anthers. (A) Mean-difference (MA) plots of PHASloci based on sRNA-seq. (B) Total abundance (mean ± SE) of meiotic 24-nt phasiRNAs in ago18 triple homozygous mutant plants (ago18-HM) and their triple heterozygous siblings (ago18-HT). Results of Student’s t test are shown in SI Appendix, Fig. S13. (C) MA plots from differential accumulation analyses of phasiRNA abundance per loci based on TraPR sRNA-seq. In A and C, average CPM values were calculated for only triple heterozygous samples. Significantly differentially expressed loci (FC > 1.5, FDR < 0.05) are represented by red circles/triangles/plus signs, and the other loci in blue. Dash lines indicate FC = ±1.5 (i.e., log2FC = ±0.585).
|
|
|
|
[ Other News ]___________________________________________________
|


 Curently online :
Curently online :
 Total visitors :
Total visitors :
(82).png)