Sugarcane, the world's most harvested crop by tonnage, has shaped global history, trade and geopolitics, and is currently responsible for 80% of sugar production worldwide1. While traditional sugarcane breeding methods have effectively generated cultivars adapted to new environments and pathogens, sugar yield improvements have recently plateaued2.
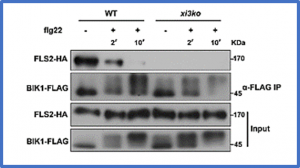
Plants rely on immune receptor complexes at the cell surface to perceive microbial molecules and transduce these signals into the cell to regulate immunity. Various immune receptors and associated proteins are often dynamically distributed in specific nanodomains on the plasma membrane (PM). However, the exact molecular mechanism and functional relevance of this nanodomain targeting in plant immunity regulation remain largely unknown.
Temperate Geng/Japonica (GJ) rice yields have improved significantly, bolstering global food security. However, GJ rice breeding faces challenges, including enhancing grain quality, ensuring stable yields at warmer temperatures, and utilizing alkaline land. In this study, we employed CRISPR/Cas9 gene-editing technology to knock out the GS3 locus in seven elite GJ varieties with superior yield performance.
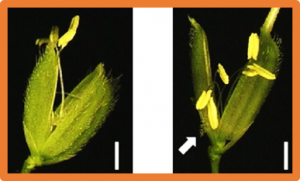
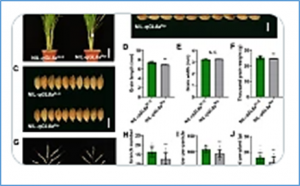
Grain size is a crucial agronomic trait that determines grain weight and final yield. Although several genes have been reported to regulate grain size in rice (Oryza sativa), the function of Wall-Associated Kinase family genes affecting grain size is still largely unknown. In this study, we identified GRAIN WEIGHT AND NUMBER 1 (GWN1) using map-based cloning. GWN1 encodes the OsWAK74 protein kinase, which is conserved in plants.
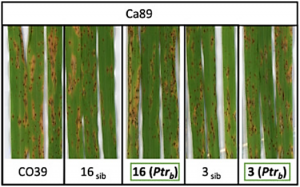
A critical step to maximize the usefulness of genome-wide association studies (GWAS) in plant breeding is the identification and validation of candidate genes underlying genetic associations. This is of particular importance in disease resistance breeding where allelic variants of resistance genes often confer resistance to distinct populations, or races, of a pathogen. Here, we perform a genome-wide association analysis of rice blast resistance in 500 genetically diverse rice accessions. To facilitate candidate gene identification, we produce de-novo genome assemblies of ten rice accessions with various rice blast resistance associations.
Endoplasmic reticulum (ER)–associated degradation (ERAD) plays key roles in controlling protein levels and quality in eukaryotes. The Ring Finger Protein 185 (RNF185)/membralin ubiquitin ligase complex was recently identified as a branch in mammals and is essential for neuronal function, but its function in plant development is unknown. Here, we report the map-based cloning and characterization of Narrow Leaf and Dwarfism 1 (NLD1)
Increasing planting density is a key strategy to enhance maize yields1-3. An ideotype for dense planting requires a ‘smart canopy’ with leaf angles at different canopy layers differentially optimized to maximize light interception and photosynthesis4-6, amongst other features. Here, we identified leaf angle architecture of smart canopy 1 (lac1), a natural mutant possessing upright upper leaves, less erect middle leaves and relatively flat lower leaves.
Seed dormancy in wild cowpea may be useful in breeding cultivated cowpea with pre-harvest sprouting resistance. A previous study identified a major quantitative trait locus (QTL) for seed dormancy, Sdp1.1+ , using the population of the cross between cultivated cowpea ‘JP81610’ and wild cowpea ‘JP89083.’ However, the molecular basis of seed dormancy in cowpea is not yet known. In this study, we aimed to finely map the locus Sdp1.1+ and identify candidate gene(s) for it.
Blast disease caused by the fungus Magnaporthe oryzae is one of the most devastating rice diseases. Disease resistance genes such as Pi-ta or Pi-ta2 are critical in protecting rice production from blast. Published work reports that Pi-ta codes for a nucleotide-binding and leucine-rich repeat domain protein (NLR) that recognizes the fungal protease-like effector AVR-Pita by direct binding.
The identification of drought stress regulatory genes is essential for the genetic improvement of maize yield. Deoxyribonucleic acid binding with one finger (Dof), a plant-specific transcription factor family, is involved in signal transduction, morphogenesis, and environmental stress responses. In present study, by weighted correlation network analysis (WGCNA) and gene co-expression network analysis, 15 putative Dof genes were identified from maize that respond to drought and rewatering.


 Curently online :
Curently online :
 Total visitors :
Total visitors :